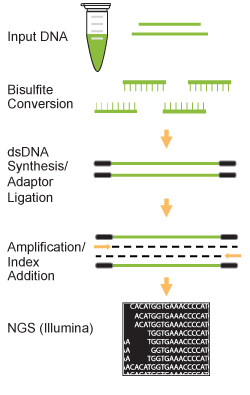
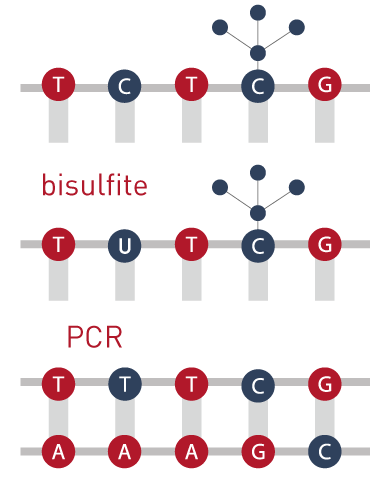
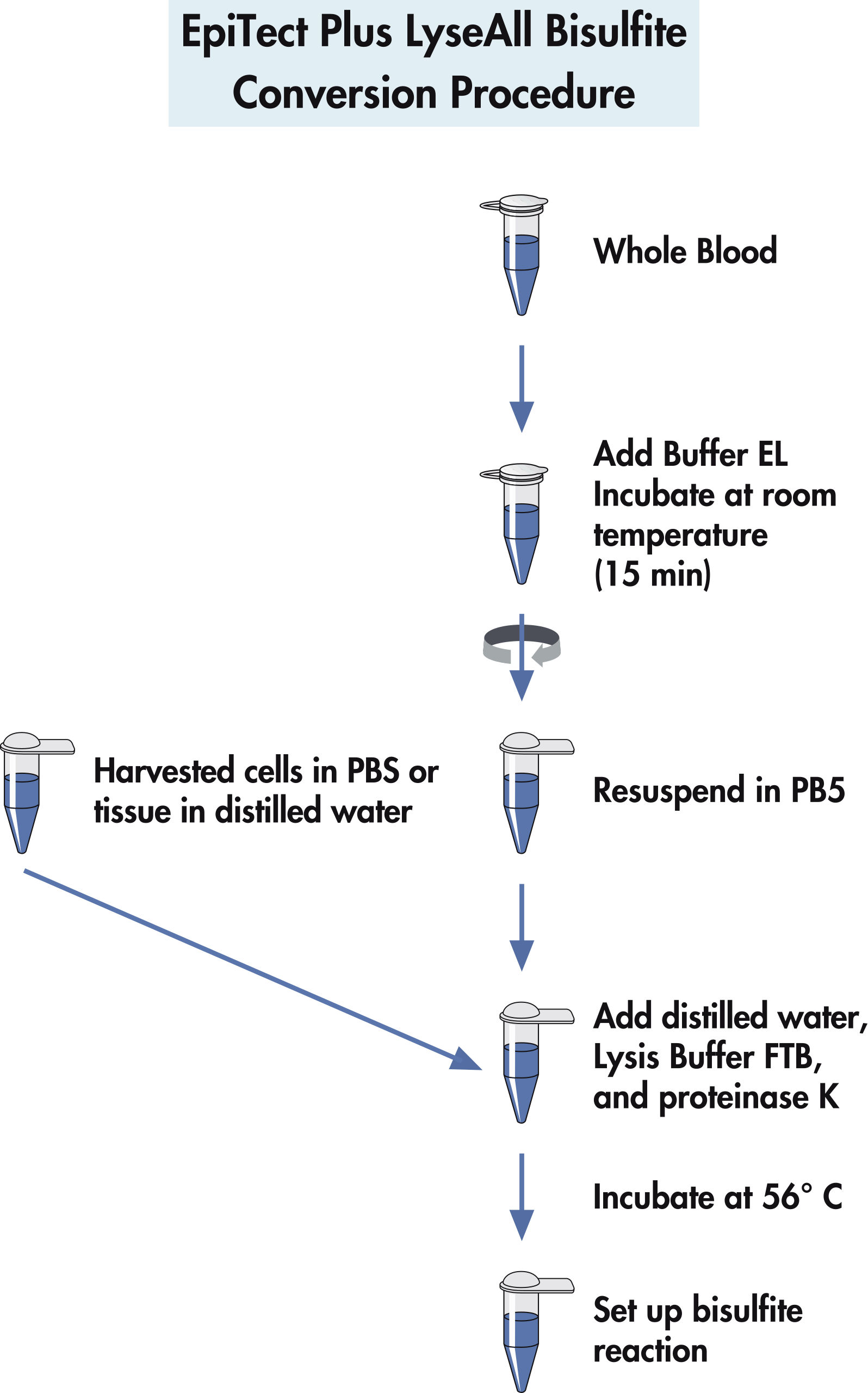
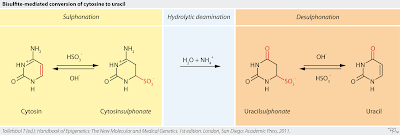
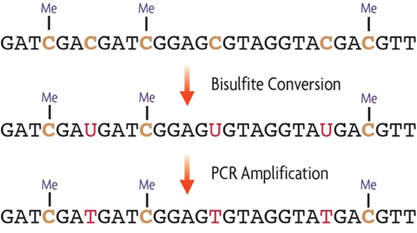
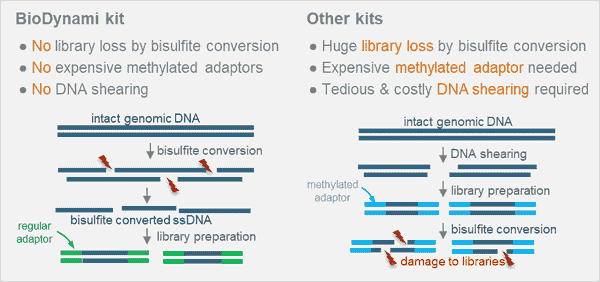
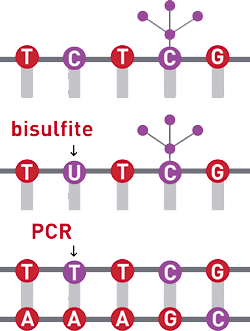
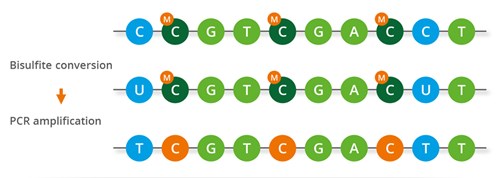
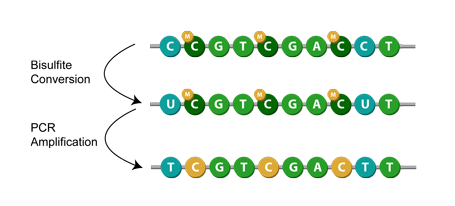Evaluation of bisulfite kits for DNA methylation profiling in terms of DNA fragmentation and DNA recovery using digital PCR | PLOS ONE
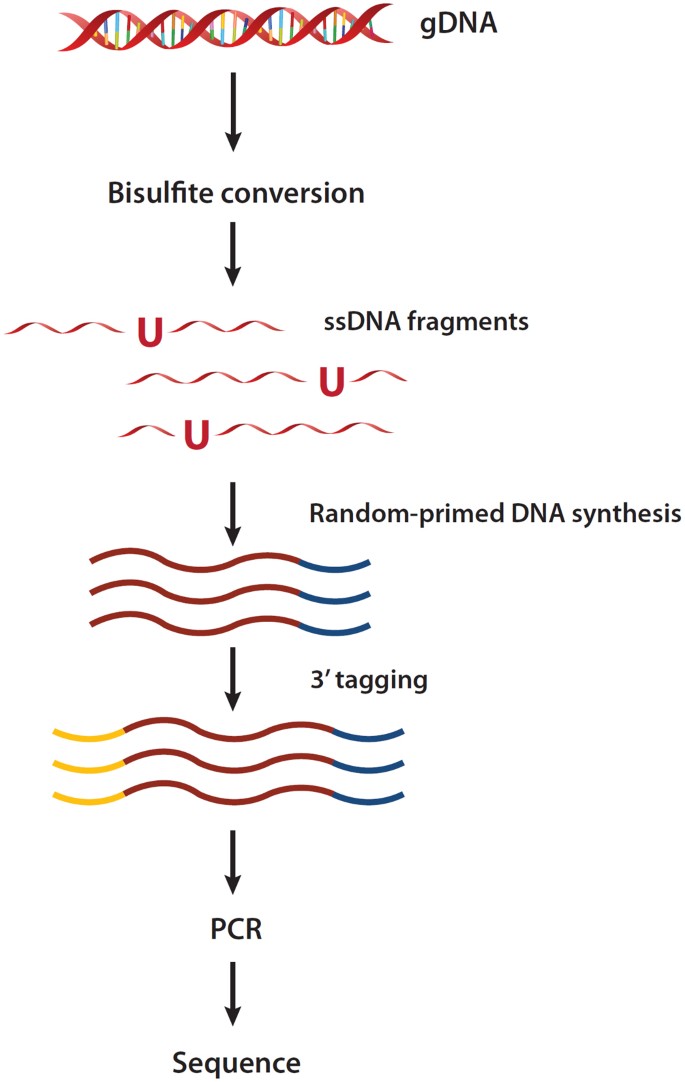
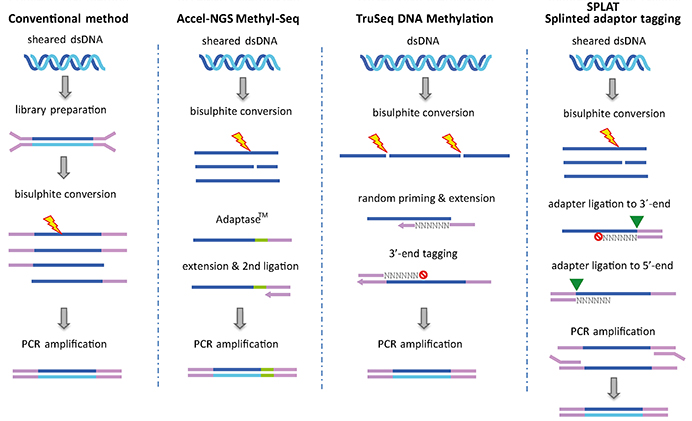
EpiGnome™ Methyl-Seq Kit: a novel post–bisulfite conversion library prep method for methylation analysis | Nature Methods

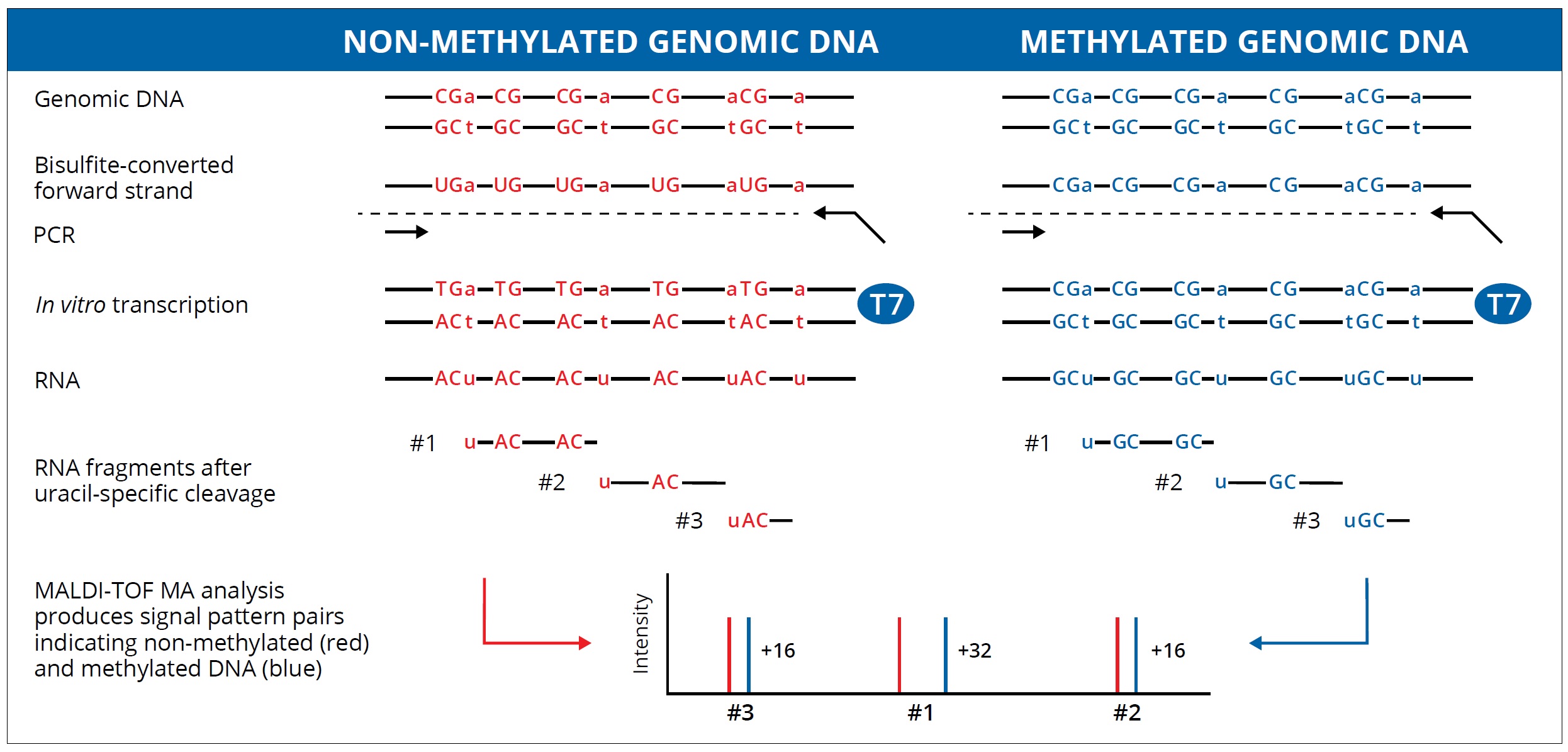
Bisulfite-converted duplexes for the strand-specific detection and quantification of rare mutations | PNAS